Introduction to Self-Incompatibility and Self-Compatibility in Plants
Self-incompatibility (SI) is a fundamental biological mechanism widely present in flowering plants, particularly in hermaphroditic species, where it functions to promote outcrossing and prevent self-fertilization. Originally described by Charles Darwin, SI refers to the ability of plants to recognize and reject their own pollen, thereby avoiding inbreeding and maintaining high levels of genetic diversity.
From an evolutionary perspective, SI is considered an ancestral reproductive strategy in angiosperms. Early flowering plants retained this mechanism to enhance cross-pollination efficiency, ensuring adaptability to diverse environmental conditions and ecological niches. This genetic system plays a critical role in population resilience, species survival, and long-term evolutionary success.
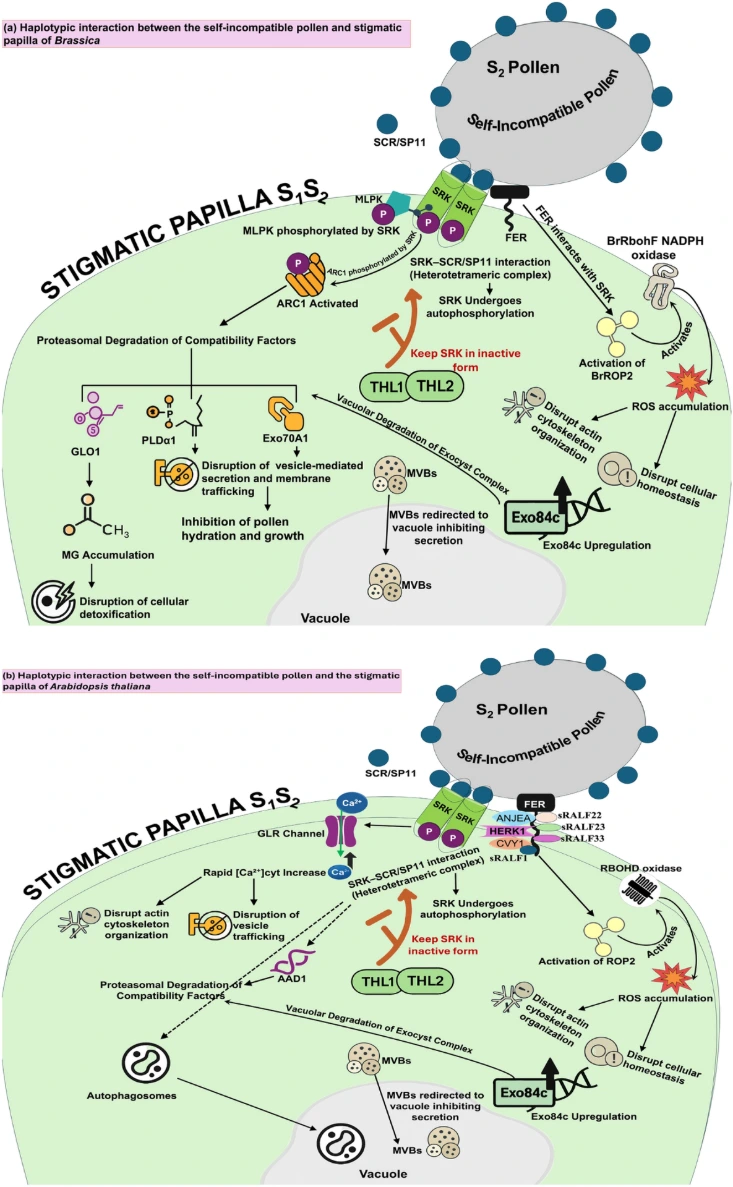
At the molecular level, SI is typically controlled by a highly polymorphic genomic region known as the S locus, which contains tightly linked genes encoding male and female determinants of compatibility. These determinants mediate a highly specific recognition system between pollen and pistil tissues. When the alleles carried by pollen and stigma match, the pollen is identified as “self” and its growth is inhibited, effectively blocking fertilization and preventing inbreeding.
Genetic and Molecular Mechanisms of Self-Incompatibility in Grasses
In angiosperms, SI systems are genetically diverse and often family-specific. They can be broadly classified into gametophytic and sporophytic systems, depending on whether pollen compatibility is determined by its own genotype or by the genotype of the parent plant.
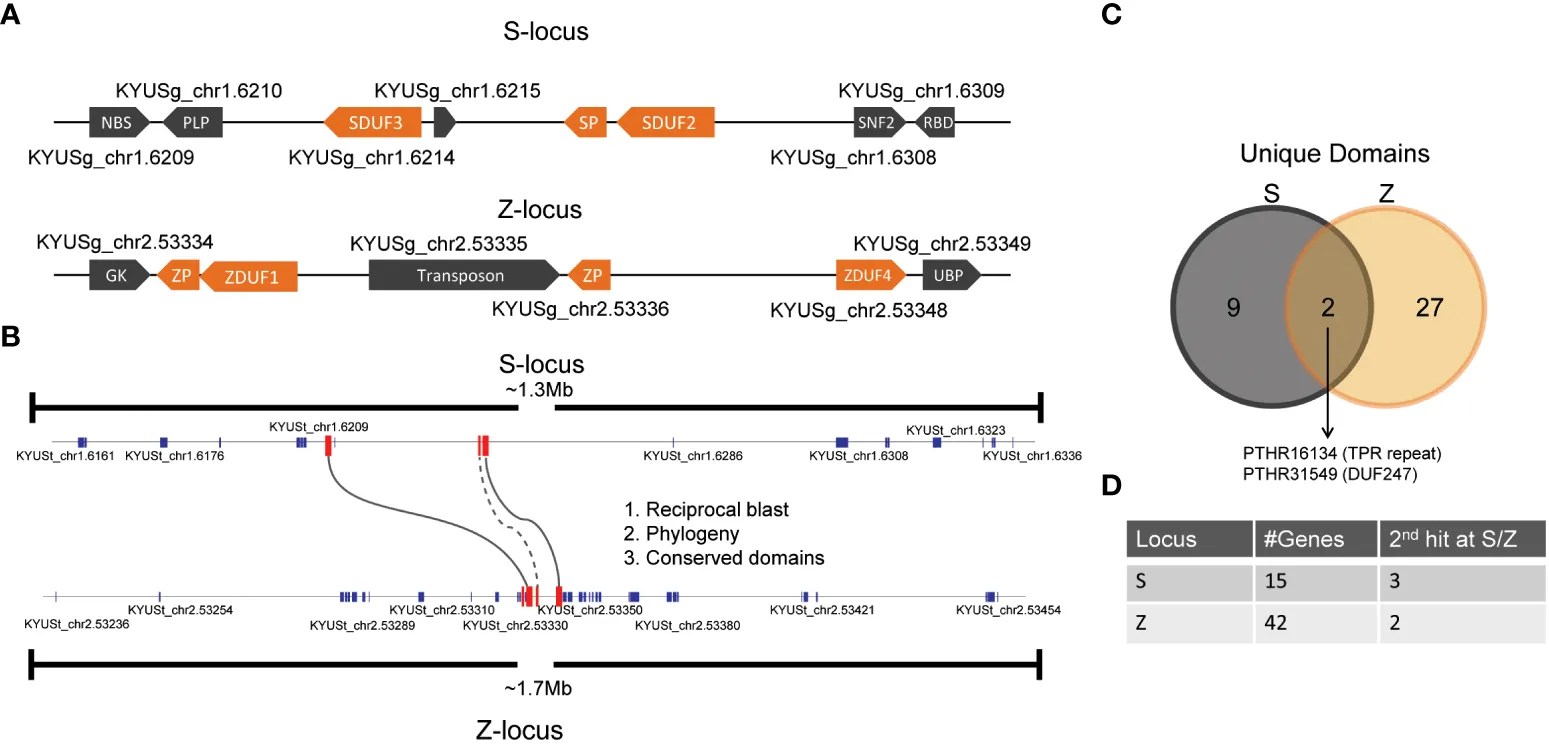
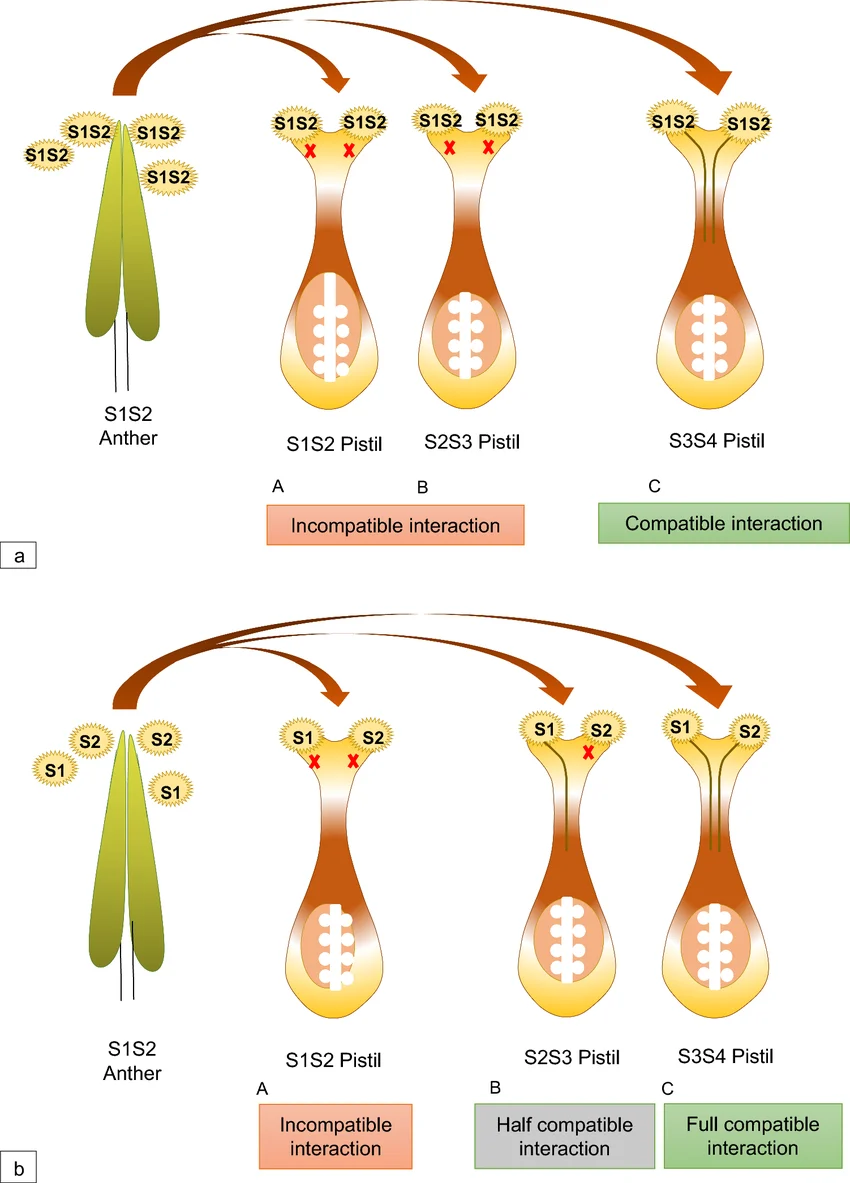
In grasses belonging to the Poaceae family, SI is controlled by a two-locus gametophytic system, involving the S and Z loci located on separate chromosomes. Fertilization is prevented when pollen and stigma share identical alleles at both loci, ensuring strict outcrossing.
Unlike single-locus systems found in other plant families, the dual-locus system in grasses adds complexity to genetic analysis and breeding applications. Although the exact identity of SI determinant genes in grasses is still under investigation, recent advances in genomics, transcriptomics, and comparative sequence analysis have identified promising candidate genes.
One such candidate includes genes encoding proteins with DUF247 domains, which are believed to play a role in pollen–stigma recognition. These proteins exhibit high allelic diversity, a characteristic typical of SI determinants due to frequency-dependent selection. Additionally, studies suggest that cellular signaling pathways involving calcium ions (Ca²⁺) and protein phosphorylation are critical components of the SI response.
Mechanisms Leading to Self-Compatibility in Grasses
Self-compatibility in grasses can arise through several genetic and environmental mechanisms:
-
Mutations in SI Determinant Genes
Loss-of-function mutations in genes located at the S or Z loci can disrupt pollen recognition, allowing self-fertilization. -
Modifier Genes Outside SI Loci
Genes unlinked to the S and Z loci can influence downstream signaling pathways, effectively bypassing the SI response. -
Epistatic Interactions
Genetic interactions between different loci can alter SI expression, leading to partial or complete self-compatibility. -
Environmental Factors (Pseudo-Self-Compatibility)
External conditions such as temperature and humidity can temporarily weaken SI responses, allowing limited self-fertilization.
These mechanisms highlight the complexity of reproductive control in grasses and emphasize the need for advanced molecular tools to fully understand SC development.
Practical Applications of Self-Compatibility in Plant Breeding
Development of Inbred Lines and Hybrid Breeding
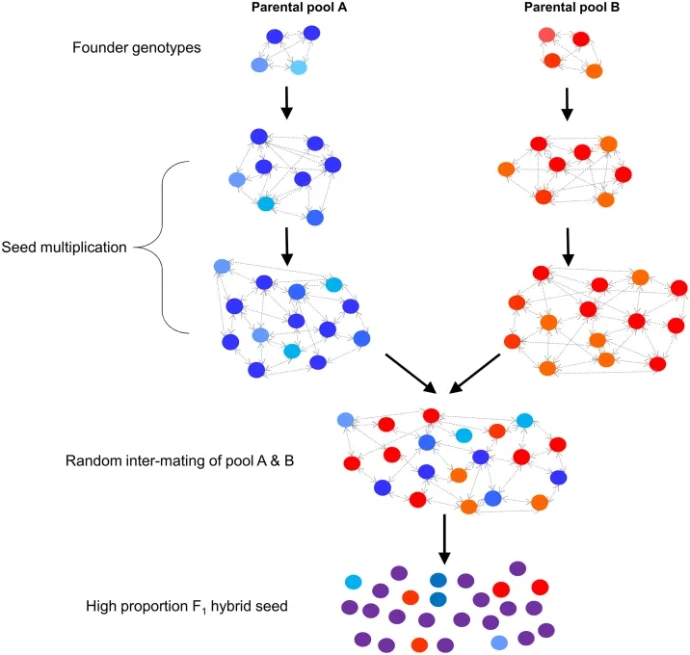
One of the most important applications of SC is the ability to produce homozygous inbred lines, which are essential for modern plant breeding. These lines allow breeders to create F1 hybrids that exhibit heterosis (hybrid vigor) a phenomenon where hybrid offspring outperform their parents in traits such as yield, growth rate, and stress tolerance.
In traditionally self-incompatible grasses, the production of inbred lines has been challenging. SC overcomes this limitation, enabling the development of stable genetic lines that can be propagated indefinitely.
Improvement of Forage Grass Breeding
Grasses such as perennial ryegrass (Lolium perenne) are essential components of global livestock production systems. However, their improvement has been limited by SI, which restricts the use of hybrid breeding techniques.
Currently, breeding programs rely on recurrent selection, which produces slow genetic gains compared to hybrid-based systems. Introducing SC into these species opens new opportunities for:
- Accelerated genetic improvement
- Enhanced biomass and seed yield
- Improved nutritional quality
- Increased resistance to diseases and environmental stress
Challenges and Future Perspectives
Despite its potential, the implementation of SC in grass breeding presents several challenges:
- Inbreeding Depression: Selfing can expose deleterious alleles, reducing plant fitness.
- Complex Genetic Control: Multiple loci and interactions complicate SC manipulation.
- Limited Germplasm Availability: SC sources are still scarce in many species.
However, ongoing advances in genomics, molecular biology, and breeding technologies are rapidly addressing these limitations. The integration of SC into breeding programs is expected to revolutionize forage crop improvement, enabling the development of high-performance, genetically stable cultivars.
Conclusion
The study and application of self-compatibility in outcrossing grass species represent a major advancement in plant genetics and agricultural biotechnology. By overcoming the limitations imposed by self-incompatibility, SC enables the development of inbred lines, hybrid breeding systems, and advanced genomic tools.
These innovations have the potential to significantly enhance crop productivity, sustainability, and resilience, particularly in economically important forage grasses. As research continues to uncover the molecular mechanisms underlying SI and SC, their practical exploitation will play a crucial role in shaping the future of plant breeding and global food security.