Introduction
The temperate grass Brachypodium distachyon has emerged as a key model organism for understanding grass biology relevant to major cereal crops such as wheat, barley, and forage grasses. With its compact genome (~272 Mb), rapid life cycle, small stature, diploid genetics, and extensive genetic resources, B. distachyon serves as a powerful system for functional genomics and trait discovery in grasses with much larger and more complex genomes.
The high‑quality genome sequence of Brachypodium provides a framework to study gene function, environmental responses, and genetic pathways that can be transferred or applied to improve agronomic traits in crops. Scientists exploit this model to identify genes controlling root development, stress responses, disease resistance, and other traits critical to sustainable agriculture and global food security.
Functional Genomics for Trait Discovery
Root traits strongly impact crop performance under water limitation and soil stress. B. distachyon has been used to simplify root research due to its genetic tractability and anatomical similarity to wheat roots, allowing identification of genes influencing root architecture. These discoveries assist in pinpointing orthologous wheat genes that may enhance water uptake and yield under limiting conditions.
Root System Architecture and Water Use Efficiency

Brachypodium distachyon diverged from wheat approximately 15 million years after the rice wheat split, making it a closer relative to wheat than rice. This closer relationship provides two key advantages: markers or primers designed from Brachypodium sequences are more likely to work in wheat, and orthologous genes are more likely to retain similar functions.
Despite some chromosomal rearrangements comparative studies show that Brachypodium exhibits better macro and micro collinearity with wheat than rice. Around 77% of Brachypodium genes have strong expressed sequence tag (EST) matches in wheat, and markers developed from Brachypodium sequences perform better in wheat than those from rice. Similar trends are observed with barley, supporting the use of Brachypodium as a more reliable model for wheat and temperate grasses.
Overall, Brachypodium provides a practical reference for marker development, gene discovery, and comparative genomics in wheat, despite some genome structural differences.
The first step in leveraging Brachypodium for crop improvement is to identify genes controlling the trait of interest. This can be done through several approaches:
(1) screening T-DNA mutant libraries to directly link a gene to a phenotype.
(2) analyzing natural accessions and contrasting genotypes to map quantitative trait loci (QTLs), often using Ecotilling on sequenced genomes.
(3) selecting candidate genes from literature or in silico analyses, which must then be validated through overexpression or silencing in Brachypodium.
Once a gene is identified, its wheat orthologue can be targeted to study or modify expression. If the orthologue is absent, the Brachypodium gene can be introduced directly into wheat to assess its effect. Non-transgenic approaches exploit natural variation in wheat: Brachypodium sequences help identify markers linked to traits, which can be used in mapping populations or marker-assisted selection. Additionally, sequencing wheat orthologues using NGS and pooling strategies allows rapid identification of beneficial alleles, reducing phenotyping costs and enabling the design of markers for breeding programs.
.
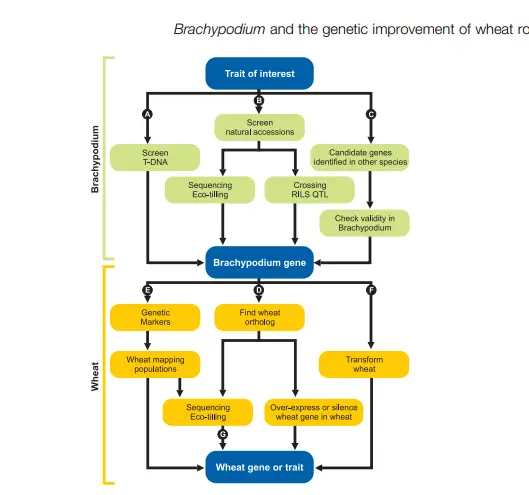 Flowchart representing the different ways to use the resources available in Brachypodium to discover new genes or traits in
wheat
Flowchart representing the different ways to use the resources available in Brachypodium to discover new genes or traits in
wheat
Stress Tolerance Mechanisms
Salt Stress Response
Salt stress reduces crop productivity by disrupting ion balance and water status. Transcriptome analysis of B. distachyon exposed to high salinity revealed thousands of differentially expressed genes, particularly those involved in oxidative stress response, osmotic regulation, and hormone‑mediated stomatal control. These gene sets serve as candidate targets for improving salt tolerance in cereal crops.
Identifying transcription factors and regulatory networks responsive to salt stress in Brachypodium provides valuable genomic loci and molecular markers that can guide genetic improvement of salt tolerance in agriculturally important species.

Transcriptional Profiling and Differential Expression Analysis
Leaves from control plants and plants treated with 200 mM NaCl for 24 h were collected for transcriptomic analysis. Total RNA was extracted using TRIzol, and RNA quality and integrity were verified using the Agilent Bioanalyzer 2100 and NanoDrop spectrophotometer. RNA-Seq libraries were prepared from 1.5 µg of total RNA using the NEBNext Ultra RNA Library Prep Kit for Illumina. mRNA was isolated using poly-T magnetic beads, fragmented, converted to cDNA, size-selected (150–200 bp), PCR-enriched, and sequenced on the Illumina platform. Experiments were performed in three biological replicates. Low-quality and adapter-containing reads were removed, and clean reads were retained for analysis.
Clean reads were aligned to the Brachypodium distachyon reference genome using HISAT2. Transcript assembly and novel transcript prediction were performed with StringTie. Gene expression levels were quantified using featureCounts and normalized as FPKM. Differential expression analysis between control and salt treated samples was conducted using DESeq2 and edgeR, with significance thresholds change.
Functional annotation and enrichment analyses were performed using GO (topGO, REVIGO) and KEGG (KOBAS), while metabolic pathway analysis was conducted with MapMan. Transcription factors were identified using the iTAK database.
To validate RNA-Seq results, quantitative real time PCR (qRT-PCR) was performed on selected upregulated and downregulated genes. cDNA was synthesized after DNase treatment, and amplification was conducted using SYBR Green chemistry on an ABI7500 system. Gene expression was normalized to the Brachypodium reference gene and calculated . Statistical significance was evaluated using Student’s t-test.
Drought Tolerance and Regulatory Genes
Drought is a major abiotic stress that impacts cereals globally. Functional characterization of B. distachyon genes such as BdWRKY36 has shown that overexpression in transgenic plants enhances drought tolerance by regulating reactive oxygen species (ROS) homeostasis and stress response pathways. Such discoveries illustrate how Brachypodium gene functions can be harnessed to improve drought resilience in target crops by identifying homologous genes or regulatory mechanisms.
Other studies involving transcriptomic and metabolomic profiling of diverse Brachypodium genotypes under drought indicate that cell wall properties and specific gene expression patterns contribute to stress tolerance and may inform breeding targets for drought‑resilient cereals.
1.Cloning and Identification of BdWRKY36
Total RNA was extracted from Brachypodium distachyon leaves and reverse-transcribed into cDNA. The BdWRKY36 open reading frame was amplified using gene-specific primers designed from genomic databases. PCR products were cloned into vectors and sequenced. Protein domain validation was performed using SMART and UniProt databases. Multiple sequence alignment and phylogenetic analysis and MEGA software to confirm gene identity and evolutionary relationships.


2.Semi-Quantitative RT-PCR
RT-PCR was used to evaluate BdWRKY36 expression across different tissues, time points, and transgenic tobacco lines. Amplification products were normalized against internal reference genes (BdUBC18 or NtUbiquitin), and transcript levels were quantified using image analysis software.

3.Gene Expression Analysis by qRT-PCR
Quantitative RT-PCR was performed to measure BdWRKY36 expression under stress conditions and to analyze stress related and ROS scavenging genes in transgenic tobacco. Specific primers were validated for efficiency and specificity. Expression levels were normalized to internal controls and calculated with three biological replicates.
4.Plasmid Construction and Subcellular Localization
The BdWRKY36 coding sequence (without stop codon) was cloned into the pCAMBIA1304 vector under the CaMV 35S promoter to generate a GFP fusion construct. The recombinant plasmid was introduced into onion epidermal cells via particle bombardment. GFP fluorescence and DAPI staining confirmed nuclear localization of the BdWRKY36 protein.


5.Transcriptional Activation in Yeast
Full-length and truncated fragments of BdWRKY36 were cloned into the pGBKT7 vector and transformed into yeast strain AH109. Growth on selective media and X-α-gal assays were used to evaluate transcriptional activation activity, confirming functional domains of the protein.
Conclusion
The Brachypodium distachyon genome has become an indispensable resource for grass genomics and crop improvement research. Its compact and well‑annotated genome allows rapid identification of genes and regulatory networks controlling traits critical for yield, stress tolerance, and disease resistance. Through functional genomics, comparative analysis, and genetic engineering tools, discoveries in Brachypodium can inform and accelerate crop improvement in wheat, barley, and other economically important cereals.
By bridging basic research with applied breeding, Brachypodium helps translate genomic knowledge into tangible advances in crop resilience, productivity, and sustainability, contributing to global food security in the face of environmental challenges.